【5.4.1】Bray-Curtis similarity index(俩群落的相似性)
一、Bray-Curtis
Bray-Curtis距离是以该统计指标的提出者J. Roger Bray和John T. Curtis的名字命名的,主要基于OTUs的计数统计,比较两个群落微生物的组成差异。与unifrac距离,包含的信息完全不一样;相比于jaccard距离,Bray-Curtis则包含了OTUs丰度信息。
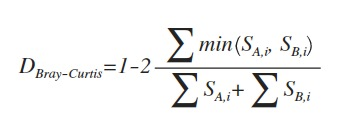
其中S_(A,i)和S_(B,i)表示第i个OTU分别在A群落和B群落中的计数。min表示取两者最小值。
举例说明
群落A和群落B的OTU统计如下表
community OTU1 OTU2 OTU3 OTU4 OTU5
A 10 8 4 1 1
B 7 3 8 4 0
min(S_(B,i)S_(B,i)) = 7+3+4+1+0 = 15
sum(S_(A,i)) = 10+8+4+1+1 = 24
sum(S_(A,i)) = 7+3+4+8+4+0 = 22
D = 1 - 2*15/(24+22) = 0.3478
当然,D值越小越好,表示二者组成差异小。
二、R的实现
vegdist {vegan}
Bray-Curtis similarity index
vegdist(x, method="bray", binary=FALSE, diag=FALSE, upper=FALSE,na.rm = FALSE, ...)
各种方法介绍:
euclidean d[jk] = sqrt(sum(x[ij]-x[ik])^2)
binary: sqrt(A+B-2*J)
manhattan d[jk] = sum(abs(x[ij] - x[ik]))
binary: A+B-2*J
gower d[jk] = (1/M) sum(abs(x[ij]-x[ik])/(max(x[i])-min(x[i])))
binary: (A+B-2*J)/M,
where M is the number of columns (excluding missing values)
altGower d[jk] = (1/NZ) sum(abs(x[ij] - x[ik]))
where NZ is the number of non-zero columns excluding double-zeros (Anderson et al. 2006).
binary: (A+B-2*J)/(A+B-J)
canberra d[jk] = (1/NZ) sum ((x[ij]-x[ik])/(x[ij]+x[ik]))
where NZ is the number of non-zero entries.
binary: (A+B-2*J)/(A+B-J)
bray d[jk] = (sum abs(x[ij]-x[ik]))/(sum (x[ij]+x[ik]))
binary: (A+B-2*J)/(A+B)
kulczynski d[jk] 1 - 0.5*((sum min(x[ij],x[ik])/(sum x[ij]) + (sum min(x[ij],x[ik])/(sum x[ik]))
binary: 1-(J/A + J/B)/2
morisita d[jk] = 1 - 2*sum(x[ij]*x[ik])/((lambda[j]+lambda[k]) * sum(x[ij])*sum(x[ik])), where
lambda[j] = sum(x[ij]*(x[ij]-1))/sum(x[ij])*sum(x[ij]-1)
binary: cannot be calculated
horn Like morisita, but lambda[j] = sum(x[ij]^2)/(sum(x[ij])^2)
binary: (A+B-2*J)/(A+B)
binomial d[jk] = sum(x[ij]*log(x[ij]/n[i]) + x[ik]*log(x[ik]/n[i]) - n[i]*log(1/2))/n[i],
where n[i] = x[ij] + x[ik]
binary: log(2)*(A+B-2*J)
cao d[jk] = (1/S) * sum(log(n[i]/2) - (x[ij]*log(x[ik]) + x[ik]*log(x[ij]))/n[i]),where S is the number of species in compared sites and n[i] = x[ij] + x[ik]
参考资料:
药企,独角兽,苏州。团队长期招人,感兴趣的都可以发邮件聊聊:tiehan@sina.cn
![]() 个人公众号,比较懒,很少更新,可以在上面提问题,如果回复不及时,可发邮件给我: tiehan@sina.cn
个人公众号,比较懒,很少更新,可以在上面提问题,如果回复不及时,可发邮件给我: tiehan@sina.cn

